Visualisation¶
Pygna offers a range of plots to visualise the results of the analysis.
GNT plots¶
GNT barplot¶
paint-dataset-stats plots the results of of a GNT test on a barplot. Pass the results table generated by one of the functions and the output figure file (png or pdf). In case you are using a SP test, pass also ‘less’ as an alternative.
usage: pygna paint-datasets-stats [-h] [-a ALTERNATIVE] table-filename output-file
positional arguments:
table-filename pygna results table
output-file figure file, use pdf or png extension
optional arguments:
-h, --help show this help message and exit
-a ALTERNATIVE, --alternative ALTERNATIVE
'greater'
Example usage: .. code-block:: bash
$ pygna paint-dataset-stats table_topology_module.csv figures/barplot_module.pdf

GNT summary¶
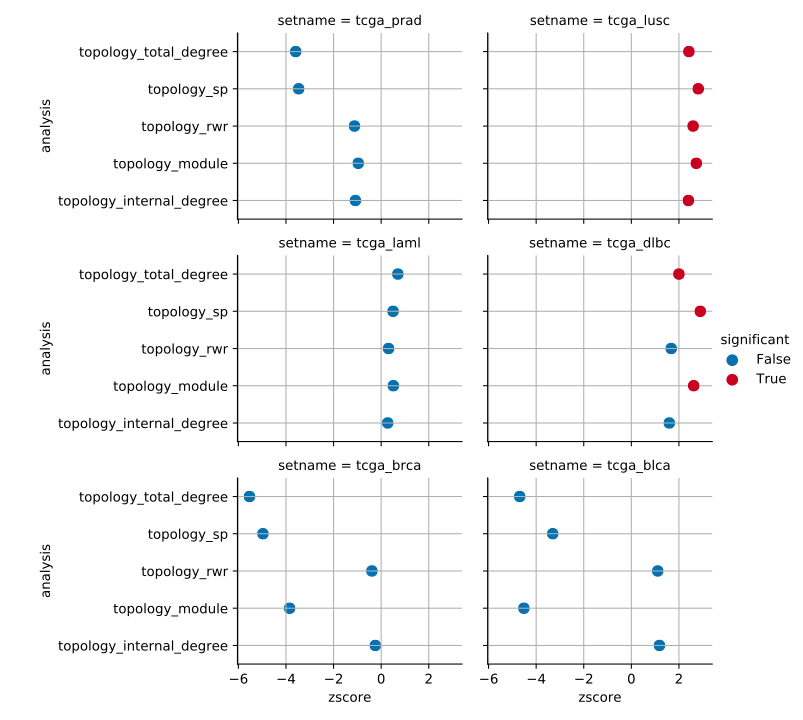
paint_summary_gnt plots a pointplot with a general summary for the GNT.
For example, given multiple GNT test results saved as table_topology_<method>.csv we could visualise all results as follows.
$ pygna paint-summary-gnt figures/summary_gnt.pdf table_topology_*
usage: pygna paint-summary-gnt [-h] [-s SETNAME] [-t THRESHOLD] [-c COLUMN_FILTER] [--larger] [--less-tests LESS_TESTS] output-figure [input_tables [input_tables ...]]
positional arguments:
output-figure output figure filename
input_tables -
optional arguments:
-h, --help show this help message and exit
-s SETNAME, --setname SETNAME
name of the dataset (default: -)
-t THRESHOLD, --threshold THRESHOLD
Value to threshold the colors (default: 0.05)
-c COLUMN_FILTER, --column-filter COLUMN_FILTER
column where the threshold is applied (default: 'empirical_pvalue')
--larger if True the threshold is set as lower limit (default: False)
--less-tests LESS_TESTS
comma separated string of the tests that are significant if lower than expected, otherwise pass empty string (default: 'topology_sp')
GNA plots¶
GNA heatmap¶
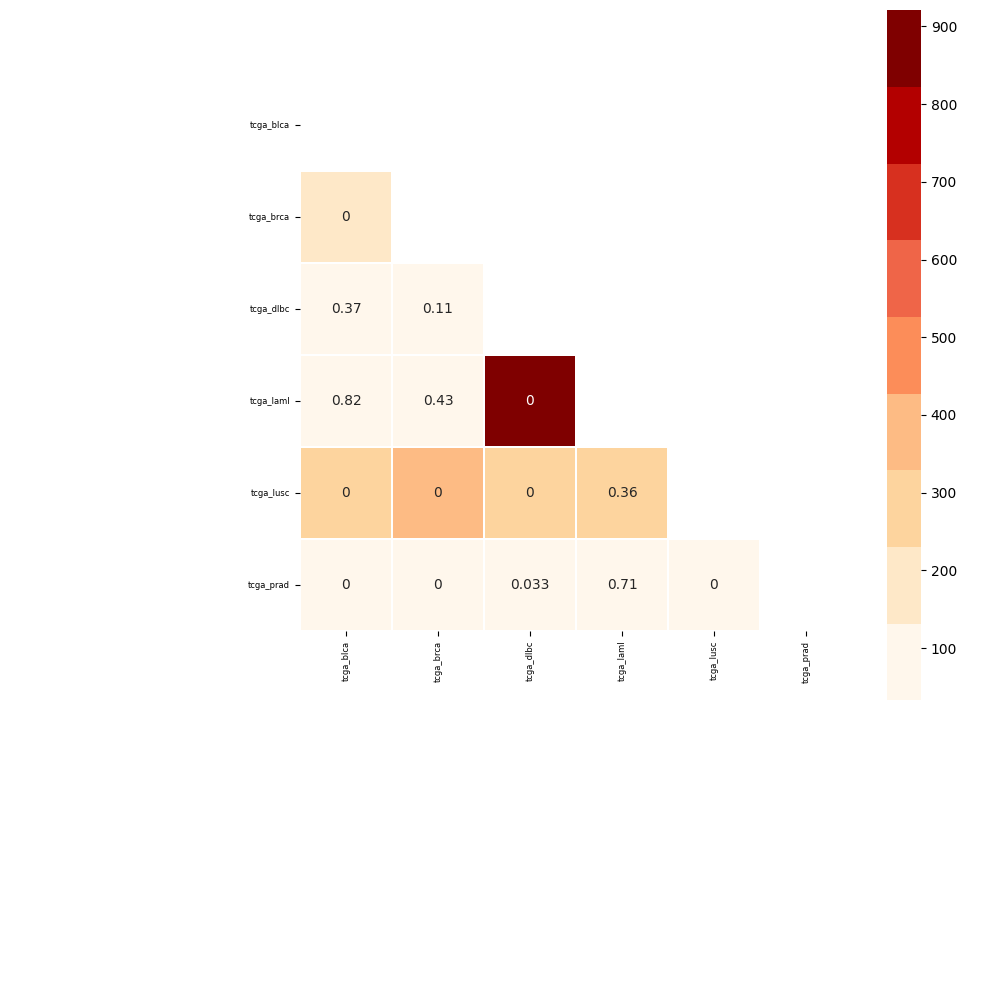
paint-comparison-matrix plots the results of of a GNA test. Pass the results table generated by one of the functions and the output figure file (png or pdf). With RWR you can specify whether the test is a RWR association, in this case a different palette and limits are sets. Specify if the results are obtained using association with only one genesets (multiple setnames in the same file). Pass the annotate flag to have the pvalue annotation on the plot
usage: pygna paint-comparison-matrix [-h] [-r] [-s] [-a] table-filename output-file
positional arguments:
table-filename pygna comparison output
output-file output figure file, specify png or pdf file
optional arguments:
-h, --help show this help message and exit
-r, --rwr use RWR is the table comes from a RWR analysis (default: False)
-s, --single-geneset use true if the comparison has been done for a single file (default: False)
-a, --annotate set true if uou want to print the pvalue inside the cell (default: False)
$ pygna paint-comparison-matrix table_comparison_rwr.csv figures/comparison_rwr.pdf -r -s -a
GNA association volcano¶
paint-volcano-plot plots the results of of a GNA test of association of a single geneset against multiple pathways. Pass the results table generated by one of the functions and the output figure file (png or pdf). From the results table, a multiple testing correction is applied and the results are those plotted.
The defined threshold are for x: zscore and \(y: -log10(pvalue)\)
usage: pygna paint-volcano-plot [-h] [-r] [-i ID_COL] [--threshold-x THRESHOLD_X] [--threshold-y THRESHOLD_Y] [-a] table-filename output-file
positional arguments:
table-filename pygna comparison output
output-file output figure file, specify png or pdf file
optional arguments:
-h, --help show this help message and exit
-r, --rwr use RWR is the table comes from a RWR analysis (default: False)
-i ID_COL, --id-col ID_COL
'setname_B'
--threshold-x THRESHOLD_X
0
--threshold-y THRESHOLD_Y
2
-a, --annotate False
